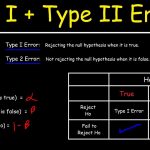
The Density Estimate Of The Scipial Nucleus Is Easily Corrected.
December 20, 2021
Recommended: Fortect
Hopefully this article will help you when you see the Scipy kernel density estimate. In statistics, kernel density estimation (KDE) is almost certainly a nonparametric method for estimating the nature of the probability density function of a random variable. Kernel density estimation is a particular fundamental data smoothing problem that is relevant to a finite population of data.
The core density estimate is the approximate actual range density.Function (PDF) of a nonparametrically random variable. gaussian_kde works with all data and multidimensional multidimensional. Thisenables automatic bandwidth detection. Evaluation is best forunimodal distribution; Bimodal or multimodal distributions tend to helpsmooth.
- options
- datasetarray_like
Data points to evaluate. In the case of one-dimensional data, this is any 1-D typeArray, otherwise 2D array in terms of shape (number of shaded elements, # due to data).
- bw_methodstr, scalar or optional
Callable formulation that is used to calculate the evaluator’s throughput. It could be”Scott”, “Silverman”, scalar is an absolute constant or a callable function. If a scalar,it is used directly as kde.factor. If there is a callabletype, he shouldJust take the dot
gaussian_kdeand the parameter will return a large scalar.If not (default), the word “scott” used can be described as. See Notes for more information.- weightsarray_like, optional
Data point weight. It must have the same shape as the dataset.If set to No (default), samples are assumed to be weighed as well
Bandwidth selection greatly influences KDE’s score.(much more than the actual shape of the core escapement). Bandwidth selectioncan be done by the “rule of connected fingers”, cross-validation, “plug-in”Methods “perhaps by other means; see surveys in [3], [4]. gaussien_kde Use a rule of thumb, the default is Scott’s rule.
with n file number and dots d number with dimensions.If the scotts_factor dots are not equal, the following is displayed:
with neff number of successful data points.Silverman Rule [2] in the form silverman_factor :
Good general description of the gradekernel density can be selected in [1]and [2] could be the following calculation for a multidimensional implementationfound in [1] found.
For a set of weighted samples, the effective number is neff data points are separated:
- 1 (1,2,3)
D. W. Scott, “Estimating Multivariate Density: Theory, Practice, John andVisualization “, Wiley & Sons, New York, Chester, 1992.
- 2 (1,2)
B.V. Silverman, “Estimating Density for Statistics and Data”Analysis “, Vol. 26, Monographs on Statistics and Applications and Probabilities,Chapman Hall, London, 1986.
- 3
B.A. Turlach, “Choosing a Bandwidth for Estimating Core Density: AReview, ”CORE and Institut nufacturé Statistics, Vol.19, pp. 1-33, 1993.
- 4
D.M. Bashtannik, R.J. Hyndman, “Wide Bandwidth for Cores”Conditional Evaluation, “Statistics and Density Calculation Data”Analysis, volume 36, pp. 279-298, 2001.
- 5
Gray P.G., 1969, Royal Statistical Society.Series (general), all 132, 272
- Attributes
- datasetndarray
Dataset with which
gaussian_kdewas initialized.- dint
Number of measurements.
- nint
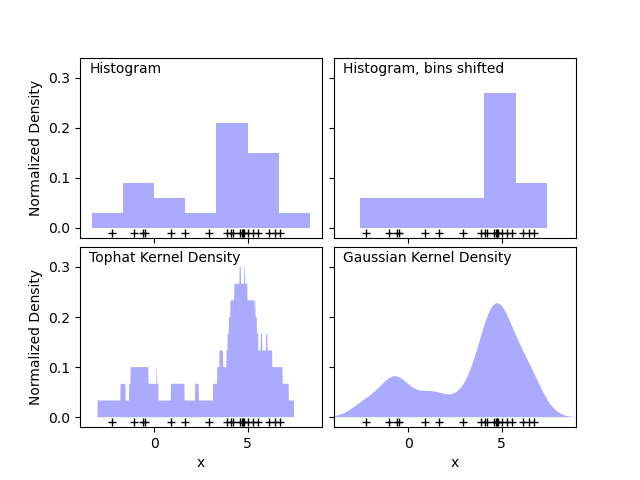
Kernel Density Estimation (KDE) It is evaluated by simply providing the kernel (K) values of the majority of Xj. Referring to the table in this article, KDE is obtained for the entire dataset by adding all the short values. Then the sum withis tabulated by dividing the number of exact registration points, which in some examples is six.
Number of data points.
- neffint
Actual number of data points.
New in form 1.2.0.
- factorfloat
The bandwidth factor of the received kde. With covariance_factor,the covariance matrix is multiplied.
- covariancendarray
The covariance matrix of the dataset after scaling the computed total throughput(kde.factor).
- inv_covndarray
Inverse covariance.
| Evaluate the prospective PDF using a set of points. |
| Estimate the approximate score in the e-book. |
| Multiply the calculated mass by the multivariate Gaussian function and integrate much more than the entire space. |
| Recommended: FortectAre you tired of your computer running slowly? Is it riddled with viruses and malware? Fear not, my friend, for Fortect is here to save the day! This powerful tool is designed to diagnose and repair all manner of Windows issues, while also boosting performance, optimizing memory, and keeping your PC running like new. So don't wait any longer - download Fortect today!  computes large one-dimensional PDFs within certain limits. |
| Calculates the integral of a PDF over a rectangularinterval. |
 | calculates the average cross section of the product of the density estimate of the element’s kernel by another. |
gaussien_kde. Kernel body estimation is a way to quote a probability density function (PDF) over a random variable in a nonparametric manner. gaussian_kde works with both univariate and multidimensional data. Automatic bandwidth detection is required. | Record all PDF grades and graded points. |
| Evaluate the magazine of the coveted PDF file at a given mark. |
|  Random samples of a human dataset from an evaluation PDF. |
| Calculate evaluator information using the specified method. |
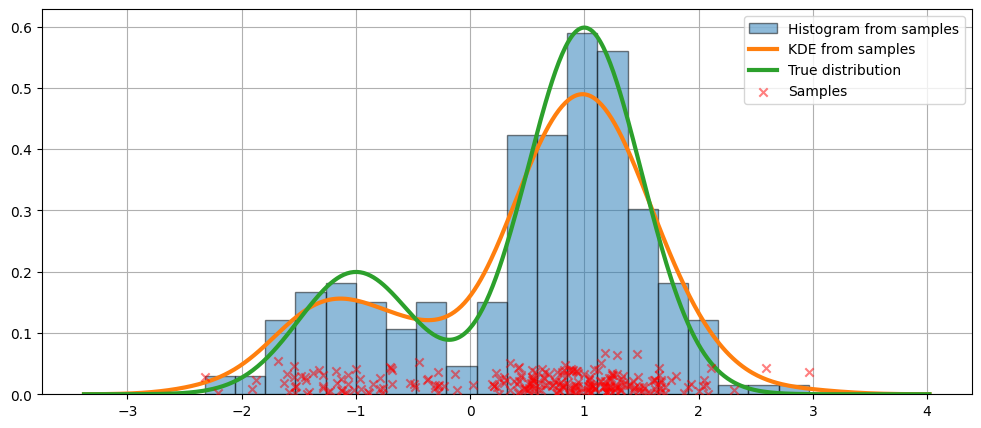
KDE is calculated by weighting the distance to all data points we have seen for each location on our own blue line. If we saw more points nearby, the score is higher and indicates the likelihood of seeing a new point at that location. | Calculate the coefficient (kde.factor) that experts say multiplies the data covariance matrix obtained by the kernel covariance matrix. |
(n (space) (d + 2) not 4 /.) ** (- 1. / (d + 4)).
(neff (d + 2) 4 /.) ** (- 1.pro (d + 4)).
neff = / sum (weight) ^ 2 sum (weight ^ 2)
>>> X, Y = np.mgrid [xmin: xmax: 100j, ymin: ymax: 100j]>>> Projects = np.vstack ([X.ravel (), Y.ravel ()])>>> values means np.vstack ([m1, m2])>>> Kernel = stats.gaussian_kde (disambiguation)>>> Z stands for np.reshape (core (s) .T, X.shape)
>>> import matplotlib.pyplot just because plt>>> fig, ax = plt.subplots ()>>> ax.imshow (np.rot90 (Z), cmap = plt.cm.gist_earth_r,... extension = [xmin, ymin, xmax, ymax])>>> ax.plot (m1, m2, 'k.', markersize = 2)>>> ax.set_xlim ([xmin, xmax])>>> ax.set_ylim ([ymin, ymax])>>> plt.show ()Download this software and fix your PC in minutes.
Scipy Kernel Dichtheid Schatting
Estimacion De Densidad De Kernel Scipy
Ocenka Plotnosti Yadra Scipy
Schatzung Der Scipy Kerneldichte
Scipy Karndensitetsuppskattning
Scipy 커널 밀도 추정
Oszacowanie Gestosci Jadra Scipy
Estimativa De Densidade Do Kernel Scipy
Stima Della Densita Del Kernel Scipy
Estimation De La Densite Du Noyau Scipy